The paper ‘Modeling the microRNA Regulation of TGF-β/SMAD Signaling Pathways for Seizure Control in Temporal Lobe Epilepsy’ written by Walton Institute at South East Technological University’s Kurt J. A. Pumares supervised by Dr Deirdre Kilbane (Director of Research, Walton Institute), in collaboration with Daniel P. Martins (University of Essex), Aiman Khalil (Walton Institute), and Jochen H. M. Prehn (Royal College of Surgeons in Ireland) was recently published in the journal npj Systems Biology and Applications.
Summary
Temporal lobe epilepsy is one of the most common types of epilepsy. Recent studies on analysing the molecular components of neurons in the brain provided potential therapeutic treatments for neurological disorders. Previously, seizure-related biological processes were studied, wherein the concentration of specific molecules and proteins were found to be altered before and after seizure occurrences. Using prior findings, a switch between “seizure” and “anti-seizure” states was identified upon suppressing molecular activities in neurons by administering molecular inhibitors. Varying dosage strategies of molecular inhibitors and the regulatory effects of the seizure-related molecules were explored to characterise the biological responses in neurons, thus offering new ways to suppress seizures in temporal lobe epilepsy patients.
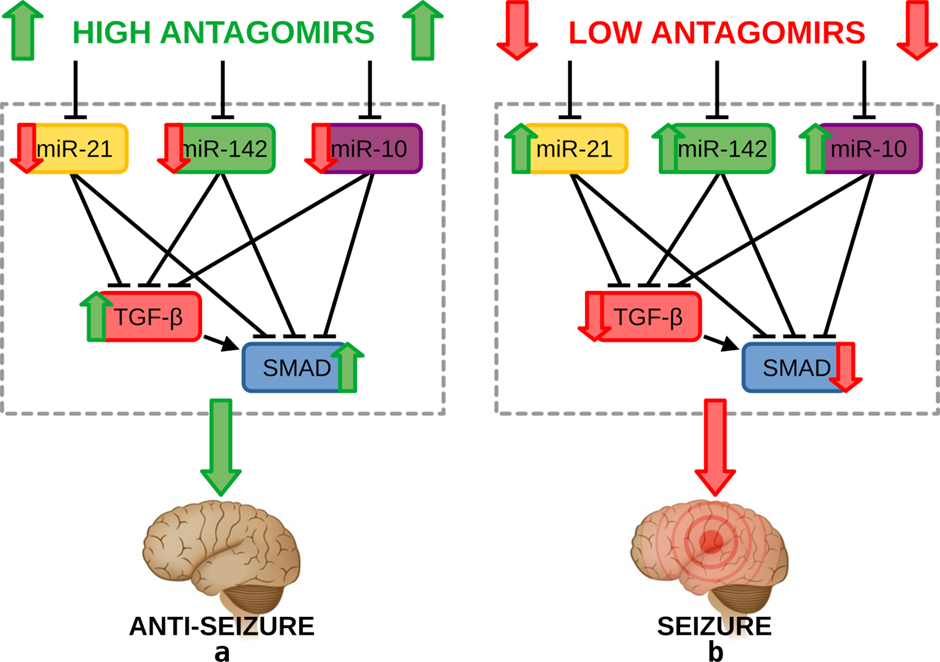
This publication outlines a mathematical model developed to characterise the seizure phenotypes based on the regulation of microRNAs – miR-21a-5p, miR-142a-5p, and miR-10a-5p – inhibiting the downstream TGF-β/SMAD signaling pathways. Strategies in administrating microRNA inhibitors called antagomirs were explored under varying dosages to validate their phenotypic switching and anti-seizure effects. These results provide insights into potential therapeutic targets for seizure suppression in temporal lobe epilepsy patients.
What does it mean for society?
Under both treatment and control strategies, the model revealed the treatment strategy to maintain the anti-seizure state while the control strategy enabled a switch into the seizure state. The individual and pairwise regulations of each microRNA were also examined and revealed varying concentration profiles for the downstream signaling pathways, thus demonstrating the effect of different model structures on phenotype switching. Moreover, bolus pre-injection, continuous infusion, and multiple dosage administration strategies were explored, suggesting various ways to promote an anti-seizure state to suppress seizures. This research altogether paves the way towards validating experimental results using modeling simulations to develop novel seizure suppression strategies through detailed analyses of intracellular signaling dynamics associated with temporal lobe epilepsy. This also provides key insights into potential therapeutic targets to develop treatment strategies for other neurological disorders.
What are the next steps?
Future works include considering other more complex modeling approaches such as exploring artificial intelligence models, particularly gene regulatory networks to reveal more useful insights and potential therapeutic targets to assist in developing seizure suppression strategies for temporal lobe epilepsy. Moreover, designing strategies to address other pathological responses in temporal lobe epilepsy could also be considered, as well as developing models using experimental data on the regulation of other microenvironmental components such as transfer RNA-derived small RNAs, and circular RNAs with their target signaling pathways. Also worth considering in the future are developing post-treatment strategies in computational models to characterise epilepsy-associated seizures.
This work was conducted as part of the EU FET Open PRIME Project which is funded from the European Union’s Horizon 2020 Research and Innovation Programme under grant agreement No 964712.
Publication Title: Modeling the microRNA Regulation of TGF-β/SMAD Signaling Pathways for Seizure Control in Temporal Lobe Epilepsy
Authors: Kurt J. A. Pumares, Daniel P. Martins, Aiman Khalil, Jochen H. M. Prehn and Dr Deirdre Kilbane (Supervisor)
Name of Journal: npj Systems Biology and Applications
Link to Publication: https://doi.org/10.1038/s41540-025-00643-6
Publication Date: 15 January 2026